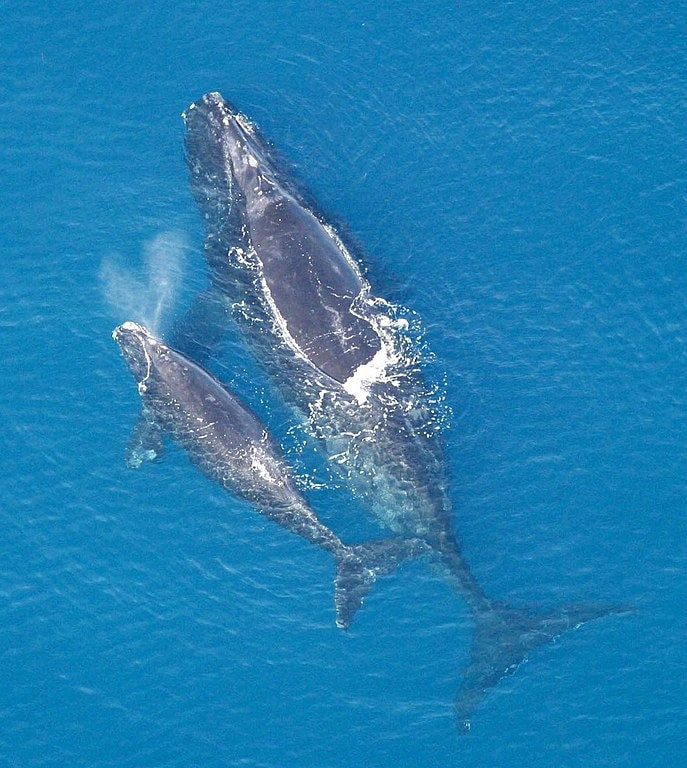
ecological, environmental & ancient genomicsconservation genomics of endangered North Atlantic right whale
ancient ancestry - Pleistocene wolf genomeS
genomic ancestry of north american canis
My post-doctoral work established the Canis Ancestry project at Princeton University and I continue to be a collaborator on this large-scale genomics project that continues to clarify ancestry of Canis species in North America. I established a RAD sequencing (RAD-seq) protocol to generate tens of thousands of genome-wide SNPs on thousands of samples from across North America. This dataset has led to the identification of genome-wide ancestry informative and adaptive markers in various Canis types that has promoted the conservation of Algonquin wolves in Canada and Red Wolves in the United States.
Read the Abstract for a recent publication on cryptic red wolf alleles in wild populations. Read the consensus publication on the uniqueness of Algonquin Wolves. Read about the discovery of red wolf ghost alleles on Galveston Island in Texas here. environmental dna
Diet Analysis with DNA Metabarcoding
Aquatic Community Biodiversity
eastern wolf survey
In 2012, I founded the Eastern Wolf Survey that focuses on Research, Conservation, and Outreach to protect the threatened Algonquin wolf. Along with my research partners, I collect noninvasive samples from wolves and coyotes to create genetic profiles that I use in Canis assignment tests. This research continues to help clarify the extent of occurrence for Eastern (Algonquin) Wolves in Ontario and Quebec. Find out more at easternwolfsurvey.ca or follow me on Twitter @EastWolfSurvey undergraduate research
other research projectsepigenetics of speciation
Although reproductive incompatibility causing speciation has been linked to variable methylation in plants, little is known about the role of methylation in differentiation leading to speciation in wild animal populations. Does variable methylation influence reproductive compatibility and to what extent does it contribute to speciation? Preliminary results are published here. california loggerhead shrike
Utilizing mtDNA sequence data and microsatellite profiles, we investigated population genetics of loggerhead shrike populations in California, including mainland California, the Channel Islands (San Clemente, Santa Catalina, Santa Cruz, & Santa Rosa), and the captive breeding population at the San Diego Zoo. Check out the publication here. |